To
Researchers
- TOP
- To Researchers
Lineup
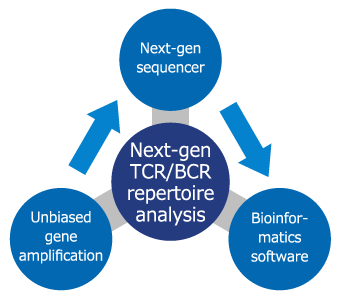
TCR/BCR Repertoire Analysis
Neoepitope Analysis
By conducting RNA seq and Exose seq together, specific gene mutations which are specific to individual cancers can be identified. In addition, by taking into account intermolecular coupling, variant peptides (neoepitope) capable of presenting an antigen will be identified. This identification can also be used for indel, long indel, and splicing variants using sequence data.
16S rRNA Bacterial Flora Analysis
Distribution of bacterial species (bacterial flora) in samples can be analyzed by examining the V1V2 or V3V4 region of the 16S rRNA gene preserved in each bacterial species using next generation sequencing. This analysis can be applied not only to biologically derived samples (feces, saliva, etc.), but also to environmentally derived samples (soil, food, etc.).
Publications
- 2024/07/22
- Paper
-
Involvement of the genus Corynebacterium in the pathogenesis of pigmented intratarsal keratinous cyst
A paper utilizing our 16S rRNA Bacterial Flora Analysis was published by Dr. Miyuki Yoshikawa, Dr. Daiki Rokunohe et al., Hirosaki University Graduate School of Medicine, Department of Dermatology, has published a paper on inflammation and bacterial flora in intratarsal keratinous cysts (IKC), which are pigmented intraocular keratinocysts.
IKC, a benign cystic lesion of the eyelid, usually appears as a yellow to white cyst, but rarely as a brown or gray-blue cyst, affecting the clinical diagnosis. The cause of this phenomenon is not known. The author's research group has observed a localized lymphocytic infiltrate in the melanocyte-rich, melanin-positive area below the cyst wall in pigmented IKC, and has analyzed the bacterial colonies within the cyst for the presence of Corynebacterium species. The etiology of pigmented IKC related to inflammation and bacterial flora was discussed in the paper.
We have performed 16S rRNA Bacterial Flora Analysis using the bacterial colonies in the cysts described in the paper.
- 2024/07/22
- Paper
-
Solid tumor
Regulatory T-cells activated in metastatic draining lymph nodes possibly suppress cancer immunity in cancer tissues of head and neck squamous cell cancerA paper utilizing our TCR Repertoire Analysis was published by Dr. Susumu Suzuki et al., Aichi Medical University Research Creation Support Center that suggests regulatory T cells activated in metastatic lymph nodes suppress cancer immunity in head and neck squamous cell carcinoma (HNSCC) cancer tissue.
Since the mechanism by which activated regulatory T cells (Tregs) suppress cancer immunity is unknown, to elucidate this mechanism, the research group performed T cell receptor (TCR) repertoire analysis was performed. We found that the TCR repertoires were biased in cancer tissue and metastatic DLNs (M-DLNs) compared to non-metastatic DLNs, and that the TCR repertoires of Tregs and CD8+ T cells between M-DLNs and cancer tissue were more similar than in other sites. These results suggest that Treg and CD8+ T cells are activated by cancer antigens such as neoantigens and shared antigens in M-DLNs and cancer tissues, and that Tregs suppress CD8+ T cell function in a cancer antigen-specific manner in M-DLNs and cancer tissues. Furthermore, M-DLN may be a source of Tregs and CD8+ T cells that are recruited to cancer tissues. These findings suggest that antigen-specific targeting of Tregs with M-DLN may represent a novel immunotherapeutic strategy for squamous cell carcinoma of the head and neck.
We have performed a TCR Repertoire Analysis using Tregs and conventional T cells in peripheral blood, influx regional lymph nodes (DLNs), and cancer tissues of patients with squamous cell carcinoma of the head and neck (HNSCC) as shown in the article.
- 2024/07/01
- Paper
-
Basal immunity
Human iPSC-derived CD4+ Treg-like cells engineered with chimeric antigen receptors control GvHD in a xenograft modelA paper utilizing our TCR Repertoire Analysis was published by Dr. Hisashi Yano, Shin Kaneko Laboratory, CiRA, Kyoto University.
In this paper, they report a success in producing CD4+ Treg-like cells derived from human iPS cells by inducing FOXP3 expression.
Using our TCR Repertoire Analysis, it was confirmed that cells cultured from non-T cell-derived iPS cells reconstitute various TCRα and β genes during their differentiation, constituting a polyclonal TCR repertoire.
TCR/BCR Repertoire analysis Guidebook
MUST READ! For users considering Repertoire analysis
An easy-to-understand repertoire analysis guidebook has been prepared which explains in plain language for the questions such as what is immune repertoire analysis and what kind of preparation is necessary.
Please request the guidebook from below if desired.