16S rRNA
Bacterial Flora
Analysis
- TOP
- To Researchers
- Our Technologies
- 16S rRNA Bacterial Flora Analysis
16S rRNA Bacterial Flora Analysis
The types of bacteria and their distribution are analyzed (bacterial flora analysis) by amplifying bacterial 16S rRNA gene using PCR and analyzing using next generation sequencing. The distribution of bacteria including intestinal bacteria and oral bacteria can be comprehensively evaluated.
We provide 16S rRNA bacterial flora analysis service for customers who do not have the necessary facilities, instrumentation, or analysis software in order to enable customers to easily perform bacterial flora analysis.
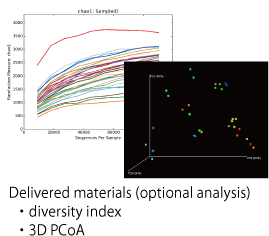
With this service, we provide a total service after receiving frozen samples from a customer including extraction of genomic DNA, 16S rRNA specific PCR, and next generation sequence analysis using Miseq as well as phylogenetic analysis and identification analysis of bacterial species using dedicated bioinformatics software (Flora Genesis). We additionally accept a second analysis (such as comparison between 2 samples, diversity index, and PCoA) according to customer’s requests.
What is 16S rRNA?
To identify types of bacteria, amplification using selective media or individual sequencing using a capillary sequencer has been necessary. However, there were issues in conducting these processes such that some bacteria not being able to be amplified or the sequencer could not be accurate for a large number of analyses. Bacterial 16S rRNA gene is an important gene involved in translation of proteins. Therefore, these genes are relatively conserved among bacterial species. Moreover, bacterial species can be identified by comparing 16s rRNA gene analysis data with databases due to the stability of the 16s rRNA gene in each bacterial species.
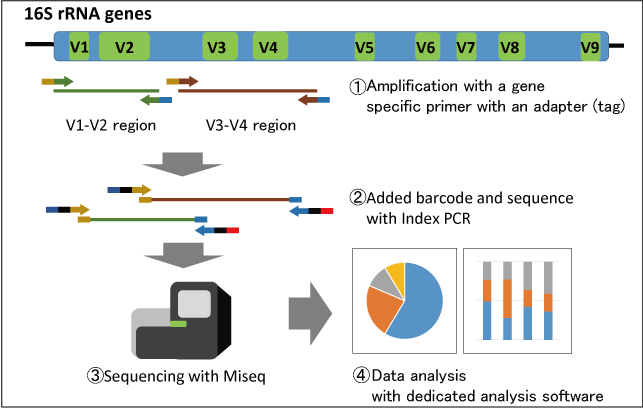
There are 9 (V1~V9) V regions in approximately 1.5kbp of the 16s rRNA gene where each V region contains a different sequence depending on the bacterial species. Approximately 500bp amplified products can be obtained by PCR using primers designed to be specific to the V1-V2 or V3-V4 regions. In fact, the genes are amplified using primers each attached to an adapter sequence for analysis using next generation sequencing followed by attaching random recognition sequence and sequence nucleotides to the products by Index PCR. This makes a sequencing possible using Miseq. The obtained sequence data are then analyzed using dedicated analysis software and primary analysis (phylogenetics analysis, bacterial flora identification analysis) of the bacterial flora can be generated. According to a customer’s request, secondary analysis (main component analysis, diversity analysis) will be performed.
Benefits of 16S rRNA Bacterial Flora Analysis
Intestinal Bacterial Flora Analysis
In recent years, it has been reported that changes in intestinal bacterial flora impact gastrointestinal immunity. Moreover, a relationship between intestinal bacterial flora and the immune system has been indicated in malignant melanoma patients. Thus, the value of the benefit of bacterial flora analysis has been demonstrated.
Analysis of Oral Icroflora
Oral bacterial flora analysis Many bacteria are known to be present in the mouth and it has been reported that such bacteria are involved not only in oral diseases but also in systemic diseases such as bacterial endocarditis and diabetes. Such diseases can be evaluated by performing oral bacterial flora analysis.
Analysis Can be Possible as Long as Samples Contain bacteria; Moreover, Samples Can be used for Various analyses
Specific samples (intravital washing solutions, surgically collected samples) which contain few bacterial flora, which is different from feces and saliva, can also be analyzed as long as they contain bacteria.Samples derived from fermented food and the environment (soils, water, air) can also be used for analysis as well as samples derived from medical materials.